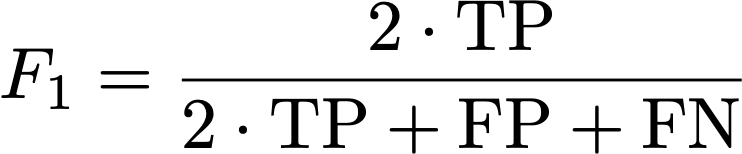Digital pathology is a fast-growing field that has seen strong scientific advances in recent years. Especially the accurate diagnosis and prognosis of tumors is a topic of particular interest, as documented by recent challenges on MICCAI, ICPR and elsewhere. In this context, the detection of cells undergoing division (mitotic figures) in histological tumor images receives high attention. The density of mitotic figures (mitotic count) is known to be related to tumor proliferation and is thus highly relevant for the prognostication. It is one of the most relevant parameters used for decision of appropriate therapy. The current gold standard method is visual assessment by a trained pathologist.
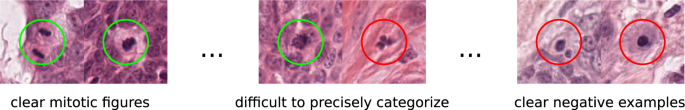
The morphology of mitotic figures strongly overlaps with similarly looking imposters. Therefore, the mitotic count is notorious for having a high inter-rater variability, which can severely impact prognostic accuracy and reproducibility. This calls for computer-augmented methods to help pathology experts in rendering a highly consistent and accurate assessment.
While previous competitions focused on the detection of mitotic figures in small sections of H&E-stained tumor tissue (e.g., the ICPR 2012 and AMIDA challenges), more recent studies moved closer to a realistic diagnostic workflow by targeting the prediction of tumor behavior on microscopy whole slide images (WSI) (e.g., the CAMELYON16/17 challenges, the PANDA challenge and MICCAI-TUPAC 16 challenge). Recent studies have shown that given a sufficient amount of high quality and high quantity annotations for tumor specimens, current deep learning-based approaches can yield performance comparable to well-trained human experts for mitotic figure identification. However, this performance severely degrades with the variability of images, caused by the tissue preparation, and image acquisition. This was the motivation for the very successful 2021 MICCAI MIDOG challenge, which is the direct predecessor of this challenge. In the past challenge, we tackled the domain shift introduced by switching to a different scanner, and the top performing approaches yielded performances in the range of human experts.
What’s different to last year (MIDOG 2021)?
Last year’s competition, however, was limited to the application on human breast cancer histopathology images. Yet, the mitotic count is of high relevance not only for these tumors, but for the diagnosis of a wide range of other neoplastic tissues as well. Between cancer tissue of different types and origin, the morphology and overall visual appearance differs strongly:
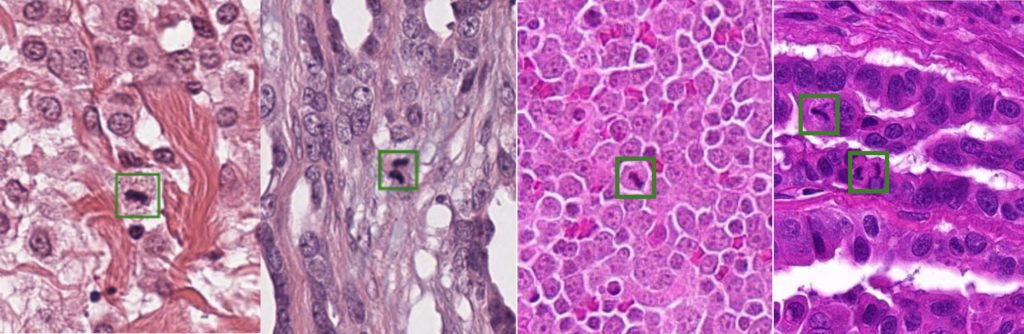
This so-called domain shift leads to a significant drop in recognition performance when naively applying machine learning algorithms trained on other tumor characteristics.
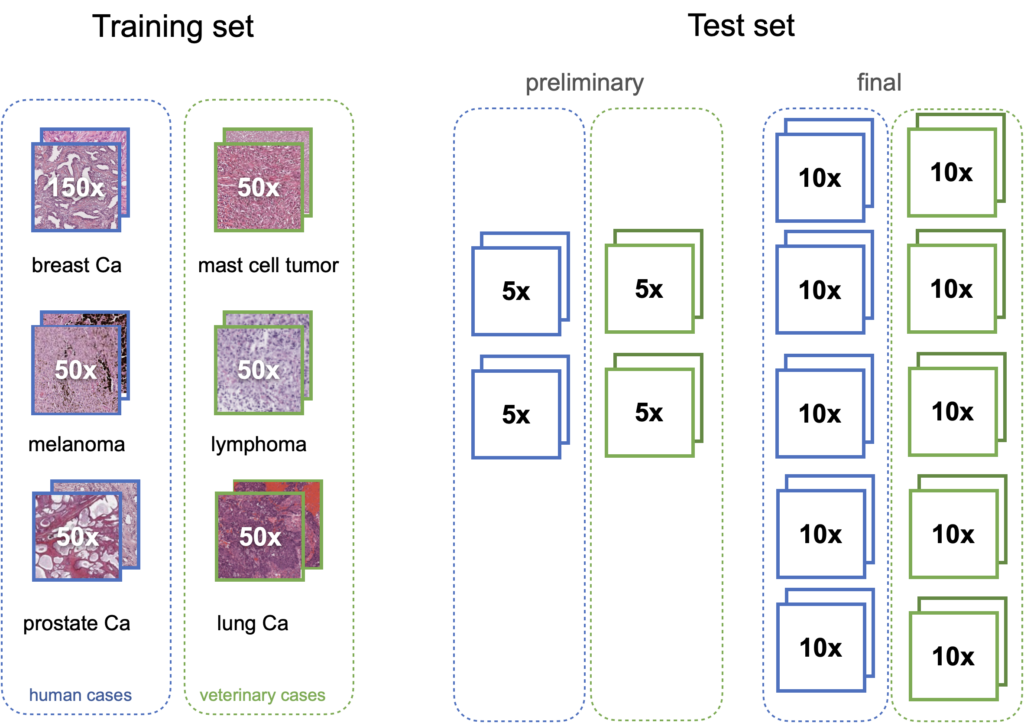
For MIDOG2, we digitize microscopy slides from 370 tumor cases, acquired from different laboratories, from different species (human, dog, cat) and with five different whole slide scanners (to reflect real-world variability). We complement these with 150 cases of human breast cancer, already available for the first MIDOG challenge, leading to 520 total cases. The training set consists of 400 cases (out of those 500) and contains at least 50 cases each of prostate carcinoma, lymphoma, lung carcinoma, melanoma, breast cancer, and mast cell tumor (six tumor types in total). The test set contains 10 cases each of 10 tumor types (100 cases in total), which are not revealed to the participants and the majority of those tumor types are unseen during training. Additionally, we have a preliminary test set for algorithm validation that contains 5 cases each of 4 different tumor types (not being part of the test set).
The target of this challenge is to develop strategies that lead to machine learning solutions that are invariant to this tissue-related domain-shift, and work equally well regardless of the tissue type that is presented to it. The challenge consists of two tasks, reflecting two different approaches towards the problem, out of which participants can select to participate in either one or both: Task 1 prohibits the use of any additional (i.e. not provided by the challenge) data and also any kind of manual or semi-automatic generation of any kind of label information. In this task, we thus expect participants to emphasize on method development regarding domain generalization. In Task 2, use of any publicly available data sets and additional labels is permitted.
Solutions that work invariant of the lab environment or the tissue have the potential to enable a whole suite of new clinical applications in tumor research and diagnostics, such as screening every tumor for missed regions of proliferation hotspots. The topic of this challenge is thus highly relevant in order to promote machine learningbased algorithms to a routine and widespread diagnostic use across laboratories.
How is it evaluated?
The common metric for evaluation mitotic figure detections is the F1 score. The main reason why we will not be using non-threshold based metrics like mAP is that the perfect detection threshold may vary strongly with the domain, so participants need to optimize a (scanner-agnostic) detection threshold.
The F1 metric accounts for false positives as well as false negatives and is thus a fair metric. As some images might have only a small number (or even none at all) mitotic figures (in low grade tumors), we calculate the F1 score over the complete data set by counting all true positives (TP), false positives (FP), and false negatives (FN) and calculate: