Training dataset
The training dataset consists of human breast cancer tissue samples acquired using four different whole slide image scanners:
- Scanner 1: Hamamatsu XR nanozoomer 2.0
- Scanner 2: Hamamatsu S360 (0.5 NA)
- Scanner 3: Aperio ScanScope CS2
- Scanner 4: Leica GT450
For each scanner, we digitized WSI of 50 different cases of breast cancer. From each WSI, a trained pathologist selected an area of 2mm² corresponding to approximately 10 high power fields, according to the grading scheme of Elston and Ellis. We cropped this area and provide it as TIFF files to ease processing.
We provide annotations for mitotic figures according to a well-established multi-expert blind annotation pipeline, aimed at finding the totality of mitotic figures (details here). We additionally provide annotations for hard examples / imposters.
Annotations are provided for scanners 1 to 3 only. The purpose of the images of scanner is to provide a reference for additional visual representation (e.g. for unsupervised domain adaptation approaches).
Distribution of mitotic count
The training set contains 1721 mitotic figures (MF) and 2714 hard examples (non-mitotic figures). The distribution of MF across scanners is rather similar:
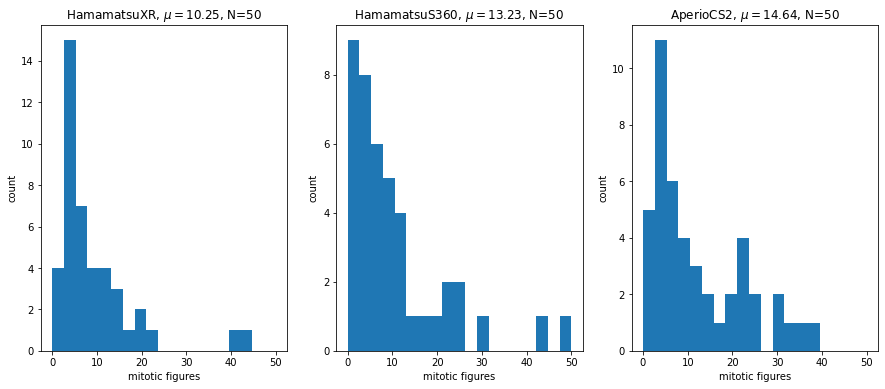
Test set
The test set contains images acquired in the same way as in the training set, but from different tumor cases. Since the challenge is about domain generalization, we only used two of the scanners that were part of the training set and added two more (undisclosed) scanners. We scanned 20 slides per scanner, corresponding to 80 cases in total.
Preliminary test set
For the self-evaluation of participating pipelines, we provide access to a preliminary test set. The preliminary test set uses the same scanners as the final test set, but consists of 5 cases per scanner (20 cases in total) only.
The access to this preliminary set is only available to the docker containers submitted to the challenge, and only available for a limited time during the competition. The purpose of this that participants can check the sanity of their approaches.
